Journal Description
BioMedInformatics
BioMedInformatics
is an international, peer-reviewed, open access journal on all areas of biomedical informatics, as well as computational biology and medicine, published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, EBSCO, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 21 days after submission; acceptance to publication is undertaken in 8.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
Latest Articles
Perspectives on Resolving Diagnostic Challenges between Myocardial Infarction and Takotsubo Cardiomyopathy Leveraging Artificial Intelligence
BioMedInformatics 2024, 4(2), 1308-1328; https://doi.org/10.3390/biomedinformatics4020072 (registering DOI) - 13 May 2024
Abstract
Background: cardiovascular diseases, including acute myocardial infarction (AMI) and takotsubo cardiomyopathy (TTC), are significant causes of morbidity and mortality worldwide. Timely differentiation of these conditions is essential for effective patient management and improved outcomes. Methods: We conducted a review focusing on studies that
[...] Read more.
Background: cardiovascular diseases, including acute myocardial infarction (AMI) and takotsubo cardiomyopathy (TTC), are significant causes of morbidity and mortality worldwide. Timely differentiation of these conditions is essential for effective patient management and improved outcomes. Methods: We conducted a review focusing on studies that applied artificial intelligence (AI) techniques to differentiate between acute myocardial infarction (AMI) and takotsubo cardiomyopathy (TTC). Inclusion criteria comprised studies utilizing various AI modalities, such as deep learning, ensemble methods, or other machine learning techniques, for discrimination between AMI and TTC. Additionally, studies employing imaging techniques, including echocardiography, cardiac magnetic resonance imaging, and coronary angiography, for cardiac disease diagnosis were considered. Publications included were limited to those available in peer-reviewed journals. Exclusion criteria were applied to studies not relevant to the discrimination between AMI and TTC, lacking detailed methodology or results pertinent to the AI application in cardiac disease diagnosis, not utilizing AI modalities or relying solely on invasive techniques for differentiation between AMI and TTC, and non-English publications. Results: The strengths and limitations of AI-based approaches are critically evaluated, including factors affecting performance, such as reliability and generalizability. The review delves into challenges associated with model interpretability, ethical implications, patient perspectives, and inconsistent image quality due to manual dependency, highlighting the need for further research. Conclusions: This review article highlights the promising advantages of AI technologies in distinguishing AMI from TTC, enabling early diagnosis and personalized treatments. However, extensive validation and real-world implementation are necessary before integrating AI tools into routine clinical practice. It is vital to emphasize that while AI can efficiently assist, it cannot entirely replace physicians. Collaborative efforts among clinicians, researchers, and AI experts are essential to unlock the potential of these transformative technologies fully.
Full article
(This article belongs to the Special Issue Computational Biology and Artificial Intelligence in Medicine)
►
Show Figures
Open AccessArticle
IMPI: An Interface for Low-Frequency Point Mutation Identification Exemplified on Resistance Mutations in Chronic Myeloid Leukemia
by
Julia Vetter, Jonathan Burghofer, Theodora Malli, Anna M. Lin, Gerald Webersinke, Markus Wiederstein, Stephan M. Winkler and Susanne Schaller
BioMedInformatics 2024, 4(2), 1289-1307; https://doi.org/10.3390/biomedinformatics4020071 (registering DOI) - 13 May 2024
Abstract
Background: In genomics, highly sensitive point mutation detection is particularly relevant for cancer diagnosis and early relapse detection. Next-generation sequencing combined with unique molecular identifiers (UMIs) is known to improve the mutation detection sensitivity. Methods: We present an open-source bioinformatics framework named Interface
[...] Read more.
Background: In genomics, highly sensitive point mutation detection is particularly relevant for cancer diagnosis and early relapse detection. Next-generation sequencing combined with unique molecular identifiers (UMIs) is known to improve the mutation detection sensitivity. Methods: We present an open-source bioinformatics framework named Interface for Point Mutation Identification (IMPI) with a graphical user interface (GUI) for processing especially small-scale NGS data to identify variants. IMPI ensures detailed UMI analysis and clustering, as well as initial raw read processing, and consensus sequence building. Furthermore, the effects of custom algorithm and parameter settings for NGS data pre-processing and UMI collapsing (e.g., UMI clustered versus unclustered (raw) reads) can be investigated. Additionally, IMPI implements optimization and quality control methods; an evolution strategy is used for parameter optimization. Results: IMPI was designed, implemented, and tested using BCR::ABL1 fusion gene kinase domain sequencing data. In summary, IMPI enables a detailed analysis of the impact of UMI clustering and parameter setting changes on the measured allele frequencies. Conclusions: Regarding the BCR::ABL1 data, IMPI’s results underlined the need for caution while designing specialized single amplicon NGS approaches due to methodical limitations (e.g., high PCR-mediated recombination rate). This cannot be corrected using UMIs.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures

Figure 1
Open AccessArticle
Cancer Classification from Gene Expression Using Ensemble Learning with an Influential Feature Selection Technique
by
Nusrath Tabassum, Md Abdus Samad Kamal, M. A. H. Akhand and Kou Yamada
BioMedInformatics 2024, 4(2), 1275-1288; https://doi.org/10.3390/biomedinformatics4020070 (registering DOI) - 13 May 2024
Abstract
Uncontrolled abnormal cell growth, known as cancer, may lead to tumors, immune system deterioration, and other fatal disability. Early cancer identification makes cancer treatment easier and increases the recovery rate, resulting in less mortality. Gene expression data play a crucial role in cancer
[...] Read more.
Uncontrolled abnormal cell growth, known as cancer, may lead to tumors, immune system deterioration, and other fatal disability. Early cancer identification makes cancer treatment easier and increases the recovery rate, resulting in less mortality. Gene expression data play a crucial role in cancer classification at an early stage. Accurate cancer classification is a complex and challenging task due to the high-dimensional nature of the gene expression data relative to the small sample size. This research proposes using a dimensionality-reduction technique to address this limitation. Specifically, the mutual information (MI) technique is first utilized to select influential biomarker genes. Next, an ensemble learning model is applied to the reduced dataset using only the most influential features (genes) to develop an effective cancer classification model. The bagging method, where the base classifiers are Multilayer Perceptrons (MLPs), is chosen as an ensemble technique. The proposed cancer classification model, the MI-Bagging method, is applied to several benchmark gene expression datasets containing distinctive cancer classes. The cancer classification accuracy of the proposed model is compared with the relevant existing methods. The experimental results indicate that the proposed model outperforms the existing methods, and it is effective and competent for cancer classification despite the limited size of gene expression data with high dimensionality. The highest accuracy achieved by the proposed method demonstrates that the proposed emerging gene-expression-based cancer classifier has the potential to help in cancer treatment and lead to a higher cancer survival rate in the future.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures

Figure 1
Open AccessArticle
A Smartphone-Based Algorithm for L Test Subtask Segmentation
by
Alexis L. McCreath Frangakis, Edward D. Lemaire and Natalie Baddour
BioMedInformatics 2024, 4(2), 1262-1274; https://doi.org/10.3390/biomedinformatics4020069 - 10 May 2024
Abstract
Background: Subtask segmentation can provide useful information from clinical tests, allowing clinicians to better assess a patient’s mobility status. A new smartphone-based algorithm was developed to segment the L Test of functional mobility into stand-up, sit-down, and turn subtasks. Methods: Twenty-one able-bodied participants
[...] Read more.
Background: Subtask segmentation can provide useful information from clinical tests, allowing clinicians to better assess a patient’s mobility status. A new smartphone-based algorithm was developed to segment the L Test of functional mobility into stand-up, sit-down, and turn subtasks. Methods: Twenty-one able-bodied participants each completed five L Test trials, with a smartphone attached to their posterior pelvis. The smartphone used a custom-designed application that collected linear acceleration, gyroscope, and magnetometer data, which were then put into a threshold-based algorithm for subtask segmentation. Results: The algorithm produced good results (>97% accuracy, >98% specificity, >74% sensitivity) for all subtasks. Conclusions: These results were a substantial improvement compared with previously published results for the L Test, as well as similar functional mobility tests. This smartphone-based approach is an accessible method for providing useful metrics from the L Test that can lead to better clinical decision-making.
Full article
(This article belongs to the Special Issue Editor's Choices Series for Methods in Biomedical Informatics Section)
►▼
Show Figures

Figure 1
Open AccessArticle
ConsensusPrime—A Bioinformatic Pipeline for Efficient Consensus Primer Design—Detection of Various Resistance and Virulence Factors in MRSA—A Case Study
by
Maximilian Collatz, Martin Reinicke, Celia Diezel, Sascha D. Braun, Stefan Monecke, Annett Reissig and Ralf Ehricht
BioMedInformatics 2024, 4(2), 1249-1261; https://doi.org/10.3390/biomedinformatics4020068 - 10 May 2024
Abstract
Background: The effectiveness and reliability of diagnostic tests that detect DNA sequences largely hinge on the quality of the used primers and probes. This importance is especially evident when considering the specific sample being analyzed, as it affects the molecular background and potential
[...] Read more.
Background: The effectiveness and reliability of diagnostic tests that detect DNA sequences largely hinge on the quality of the used primers and probes. This importance is especially evident when considering the specific sample being analyzed, as it affects the molecular background and potential for cross-reactivity, ultimately determining the test’s performance. Methods: Predicting primers based on the consensus sequence of the target has multiple advantages, including high specificity, diagnostic reliability, broad applicability, and long-term validity. Automated curation of the input sequences ensures high-quality primers and probes. Results: Here, we present a use case for developing a set of consensus primers and probes to identify antibiotic resistance and virulence genes in Staphylococcus (S.) aureus using the ConsensusPrime pipeline. Extensive qPCR experiments with several S. aureus strains confirm the exceptional quality of the primers designed using the pipeline. Conclusions: By improving the quality of the input sequences and using the consensus sequence as a basis, the ConsensusPrime pipeline pipeline ensures high-quality primers and probes, which should be the basis of molecular assays.
Full article
(This article belongs to the Special Issue Editor's Choice Series for the Computational Biology and Medicine Section)
►▼
Show Figures

Figure 1
Open AccessReview
Current Applications of Artificial Intelligence in the Neonatal Intensive Care Unit
by
Dimitrios Rallis, Maria Baltogianni, Konstantina Kapetaniou and Vasileios Giapros
BioMedInformatics 2024, 4(2), 1225-1248; https://doi.org/10.3390/biomedinformatics4020067 - 9 May 2024
Abstract
Artificial intelligence (AI) refers to computer algorithms that replicate the cognitive function of humans. Machine learning is widely applicable using structured and unstructured data, while deep learning is derived from the neural networks of the human brain that process and interpret information. During
[...] Read more.
Artificial intelligence (AI) refers to computer algorithms that replicate the cognitive function of humans. Machine learning is widely applicable using structured and unstructured data, while deep learning is derived from the neural networks of the human brain that process and interpret information. During the last decades, AI has been introduced in several aspects of healthcare. In this review, we aim to present the current application of AI in the neonatal intensive care unit. AI-based models have been applied to neurocritical care, including automated seizure detection algorithms and electroencephalogram-based hypoxic-ischemic encephalopathy severity grading systems. Moreover, AI models evaluating magnetic resonance imaging contributed to the progress of the evaluation of the neonatal developing brain and the understanding of how prenatal events affect both structural and functional network topologies. Furthermore, AI algorithms have been applied to predict the development of bronchopulmonary dysplasia and assess the extubation readiness of preterm neonates. Automated models have been also used for the detection of retinopathy of prematurity and the need for treatment. Among others, AI algorithms have been utilized for the detection of sepsis, the need for patent ductus arteriosus treatment, the evaluation of jaundice, and the detection of gastrointestinal morbidities. Finally, AI prediction models have been constructed for the evaluation of the neurodevelopmental outcome and the overall mortality of neonates. Although the application of AI in neonatology is encouraging, further research in AI models is warranted in the future including retraining clinical trials, validating the outcomes, and addressing serious ethics issues.
Full article
(This article belongs to the Special Issue Editor-in-Chief's Choices in Biomedical Informatics)
►▼
Show Figures

Figure 1
Open AccessArticle
Selection of the Discriming Feature Using the BEMD’s BIMF for Classification of Breast Cancer Mammography Image
by
Fatima Ghazi, Aziza Benkuider, Fouad Ayoub and Khalil Ibrahimi
BioMedInformatics 2024, 4(2), 1202-1224; https://doi.org/10.3390/biomedinformatics4020066 - 9 May 2024
Abstract
Mammogram exam images are useful in identifying diseases, such as breast cancer, which is one of the deadliest cancers, affecting adult women around the world. Computational image analysis and machine learning techniques can help experts identify abnormalities in these images. In this work
[...] Read more.
Mammogram exam images are useful in identifying diseases, such as breast cancer, which is one of the deadliest cancers, affecting adult women around the world. Computational image analysis and machine learning techniques can help experts identify abnormalities in these images. In this work we present a new system to help diagnose and analyze breast mammogram images. To do this, the system a method the Selection of the Most Discriminant Attributes of the images preprocessed by BEMD “SMDA-BEMD”, this entails picking the most pertinent traits from the collection of variables that characterize the state under study. A reduction of attribute based on a transformation of the data also called an extraction of characteristics by extracting the Haralick attributes from the Co-occurrence Matrices Methods “GLCM” this reduction which consists of replacing the initial set of data by a new reduced set, constructed at from the initial set of features extracted by images decomposed using Bidimensional Empirical Multimodal Decomposition “BEMD”, for discrimination of breast mammogram images (healthy and pathology) using BEMD. This decomposition makes it possible to decompose an image into several Bidimensional Intrinsic Mode Functions “BIMFs” modes and a residue. The results obtained show that mammographic images can be represented in a relatively short space by selecting the most discriminating features based on a supervised method where they can be differentiated with high reliability between healthy mammographic images and pathologies, However, certain aspects and findings demonstrate how successful the suggested strategy is to detect the tumor. A BEMD technique is used as preprocessing on mammographic images. This suggested methodology makes it possible to obtain consistent results and establishes the discrimination threshold for mammography images (healthy and pathological), the classification rate is improved (98.6%) compared to existing cutting-edge techniques in the field. This approach is tested and validated on mammographic medical images from the Kenitra-Morocco reproductive health reference center (CRSRKM) which contains breast mammographic images of normal and pathological cases.
Full article
(This article belongs to the Special Issue Feature Papers on Methods in Biomedical Informatics)
►▼
Show Figures

Figure 1
Open AccessArticle
Diagnostic Tool for Early Detection of Rheumatic Disorders Using Machine Learning Algorithm and Predictive Models
by
Godfrey A. Mills, Dzifa Dey, Mohammed Kassim, Aminu Yiwere and Kenneth Broni
BioMedInformatics 2024, 4(2), 1174-1201; https://doi.org/10.3390/biomedinformatics4020065 (registering DOI) - 8 May 2024
Abstract
Background: Rheumatic diseases are chronic diseases that affect joints, tendons, ligaments, bones, muscles, and other vital organs. Detection of rheumatic diseases is a complex process that requires careful analysis of heterogeneous content from clinical examinations, patient history, and laboratory investigations. Machine learning techniques
[...] Read more.
Background: Rheumatic diseases are chronic diseases that affect joints, tendons, ligaments, bones, muscles, and other vital organs. Detection of rheumatic diseases is a complex process that requires careful analysis of heterogeneous content from clinical examinations, patient history, and laboratory investigations. Machine learning techniques have made it possible to integrate such techniques into the complex diagnostic process to identify inherent features that lead to disease formation, development, and progression for remedial measures. Methods: An automated diagnostic tool using a multilayer neural network computational engine is presented to detect rheumatic disorders and the type of underlying disorder for therapeutic strategies. Rheumatic disorders considered are rheumatoid arthritis, osteoarthritis, and systemic lupus erythematosus. The detection system was trained and tested using 70% and 30% respectively of labelled synthetic dataset of 100,000 records containing both single and multiple disorders. Results: The detection system was able to detect and predict underlying disorders with accuracy of 97.48%, sensitivity of 96.80%, and specificity of 97.50%. Conclusion: The good performance suggests that this solution is robust enough and can be implemented for screening patients for intervention measures. This is a much-needed solution in environments with limited specialists, as the solution promotes task-shifting from the specialist level to the primary healthcare physicians.
Full article
(This article belongs to the Special Issue Editor's Choice Series for the Applied Biomedical Data Science Section)
►▼
Show Figures

Figure 1
Open AccessReview
An Overview of Approaches and Methods for the Cognitive Workload Estimation in Human–Machine Interaction Scenarios through Wearables Sensors
by
Sabrina Iarlori, David Perpetuini, Michele Tritto, Daniela Cardone, Alessandro Tiberio, Manish Chinthakindi, Chiara Filippini, Luca Cavanini, Alessandro Freddi, Francesco Ferracuti, Arcangelo Merla and Andrea Monteriù
BioMedInformatics 2024, 4(2), 1155-1173; https://doi.org/10.3390/biomedinformatics4020064 - 7 May 2024
Abstract
Background: Human-Machine Interaction (HMI) has been an important field of research in recent years, since machines will continue to be embedded in many human actvities in several contexts, such as industry and healthcare. Monitoring in an ecological mannerthe cognitive workload (CW) of users,
[...] Read more.
Background: Human-Machine Interaction (HMI) has been an important field of research in recent years, since machines will continue to be embedded in many human actvities in several contexts, such as industry and healthcare. Monitoring in an ecological mannerthe cognitive workload (CW) of users, who interact with machines, is crucial to assess their level of engagement in activities and the required effort, with the goal of preventing stressful circumstances. This study provides a comprehensive analysis of the assessment of CW using wearable sensors in HMI. Methods: this narrative review explores several techniques and procedures for collecting physiological data through wearable sensors with the possibility to integrate these multiple physiological signals, providing a multimodal monitoring of the individuals’CW. Finally, it focuses on the impact of artificial intelligence methods in the physiological signals data analysis to provide models of the CW to be exploited in HMI. Results: the review provided a comprehensive evaluation of the wearables, physiological signals, and methods of data analysis for CW evaluation in HMI. Conclusion: the literature highlighted the feasibility of employing wearable sensors to collect physiological signals for an ecological CW monitoring in HMI scenarios. However, challenges remain in standardizing these measures across different populations and contexts.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures

Figure 1
Open AccessArticle
Assaying and Classifying T Cell Function by Cell Morphology
by
Xin Wang, Stacey M. Fernandes, Jennifer R. Brown and Lance C. Kam
BioMedInformatics 2024, 4(2), 1144-1154; https://doi.org/10.3390/biomedinformatics4020063 - 26 Apr 2024
Abstract
Immune cell function varies tremendously between individuals, posing a major challenge to emerging cellular immunotherapies. This report pursues the use of cell morphology as an indicator of high-level T cell function. Short-term spreading of T cells on planar, elastic surfaces was quantified by
[...] Read more.
Immune cell function varies tremendously between individuals, posing a major challenge to emerging cellular immunotherapies. This report pursues the use of cell morphology as an indicator of high-level T cell function. Short-term spreading of T cells on planar, elastic surfaces was quantified by 11 morphological parameters and analyzed to identify effects of both intrinsic and extrinsic factors. Our findings identified morphological features that varied between T cells isolated from healthy donors and those from patients being treated for Chronic Lymphocytic Leukemia (CLL). This approach also identified differences between cell responses to substrates of different elastic modulus. Combining multiple features through a machine learning approach such as Decision Tree or Random Forest provided an effective means for identifying whether T cells came from healthy or CLL donors. Further development of this approach could lead to a rapid assay of T cell function to guide cellular immunotherapy.
Full article
(This article belongs to the Special Issue Editor's Choices Series for Methods in Biomedical Informatics Section)
►▼
Show Figures
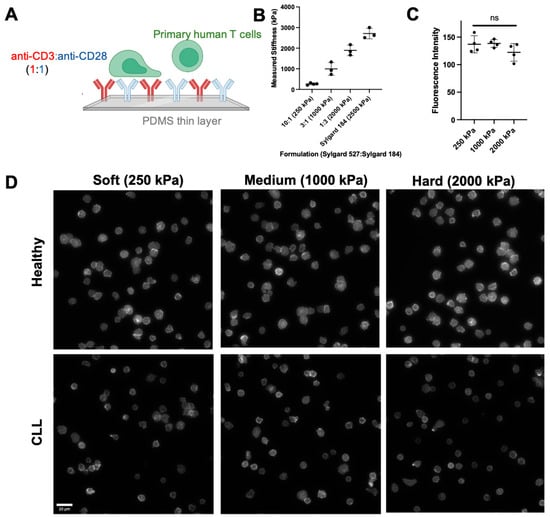
Figure 1
Open AccessReview
Recent Advances in Large Language Models for Healthcare
by
Khalid Nassiri and Moulay A. Akhloufi
BioMedInformatics 2024, 4(2), 1097-1143; https://doi.org/10.3390/biomedinformatics4020062 - 16 Apr 2024
Abstract
Recent advances in the field of large language models (LLMs) underline their high potential for applications in a variety of sectors. Their use in healthcare, in particular, holds out promising prospects for improving medical practices. As we highlight in this paper, LLMs have
[...] Read more.
Recent advances in the field of large language models (LLMs) underline their high potential for applications in a variety of sectors. Their use in healthcare, in particular, holds out promising prospects for improving medical practices. As we highlight in this paper, LLMs have demonstrated remarkable capabilities in language understanding and generation that could indeed be put to good use in the medical field. We also present the main architectures of these models, such as GPT, Bloom, or LLaMA, composed of billions of parameters. We then examine recent trends in the medical datasets used to train these models. We classify them according to different criteria, such as size, source, or subject (patient records, scientific articles, etc.). We mention that LLMs could help improve patient care, accelerate medical research, and optimize the efficiency of healthcare systems such as assisted diagnosis. We also highlight several technical and ethical issues that need to be resolved before LLMs can be used extensively in the medical field. Consequently, we propose a discussion of the capabilities offered by new generations of linguistic models and their limitations when deployed in a domain such as healthcare.
Full article
(This article belongs to the Special Issue Feature Papers in Clinical Informatics Section)
►▼
Show Figures
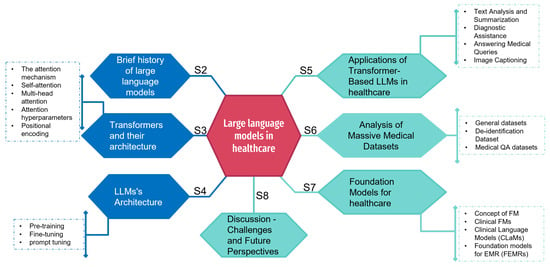
Figure 1
Open AccessProject Report
Investigating the Effectiveness of an IMU Portable Gait Analysis Device: An Application for Parkinson’s Disease Management
by
Nikos Tsotsolas, Eleni Koutsouraki, Aspasia Antonakaki, Stefanos Pizanias, Marios Kounelis, Dimitrios D. Piromalis, Dimitrios P. Kolovos, Christos Kokkotis, Themistoklis Tsatalas, George Bellis, Dimitrios Tsaopoulos, Paris Papaggelos, George Sidiropoulos and Giannis Giakas
BioMedInformatics 2024, 4(2), 1085-1096; https://doi.org/10.3390/biomedinformatics4020061 - 10 Apr 2024
Abstract
As part of two research projects, a small gait analysis device was developed for use inside and outside the home by patients themselves. The project PARMODE aims to record accurate gait measurements in patients with Parkinson’s disease (PD) and proceed with an in-depth
[...] Read more.
As part of two research projects, a small gait analysis device was developed for use inside and outside the home by patients themselves. The project PARMODE aims to record accurate gait measurements in patients with Parkinson’s disease (PD) and proceed with an in-depth analysis of the gait characteristics, while the project CPWATCHER aims to assess the quality of hand movement in cerebral palsy patients. The device was mainly developed to serve the first project with additional offline processing, including machine learning algorithms that could potentially be used for the second aim. A key feature of the device is its small size (36 mm × 46 mm × 16 mm, weight: 14 g), which was designed to meet specific requirements in terms of device consumption restrictions due to the small size of the battery and the need for autonomous operation for more than ten hours. This research work describes, on the one hand, the new device with an emphasis on its functions, and on the other hand, its connection with a web platform for reading and processing data from the devices placed on patients’ feet to record the gait characteristics of patients on a continuous basis.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
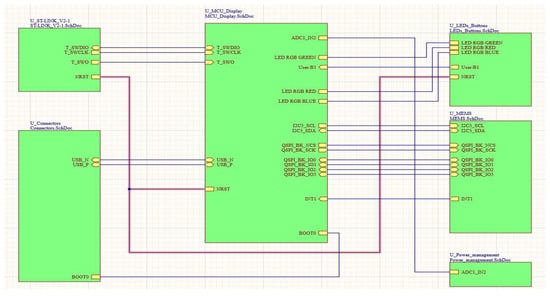
Figure 1
Open AccessArticle
Analyzing Patterns of Service Utilization Using Graph Topology to Understand the Dynamic of the Engagement of Patients with Complex Problems with Health Services
by
Jonas Bambi, Yudi Santoso, Ken Moselle, Stan Robertson, Abraham Rudnick, Ernie Chang and Alex Kuo
BioMedInformatics 2024, 4(2), 1071-1084; https://doi.org/10.3390/biomedinformatics4020060 - 9 Apr 2024
Abstract
Background: Providing care to persons with complex problems is inherently difficult due to several factors, including the impacts of proximal determinants of health, treatment response, the natural emergence of comorbidities, and service system capacity to provide timely required services. Providing visibility into the
[...] Read more.
Background: Providing care to persons with complex problems is inherently difficult due to several factors, including the impacts of proximal determinants of health, treatment response, the natural emergence of comorbidities, and service system capacity to provide timely required services. Providing visibility into the dynamics of patients’ engagement can help to optimize care for patients with complex problems. Method: In a previous work, graph machine learning and NLP methods were used to model the products of service system dynamics as atemporal entities, using a data model that collapsed patient encounter events across time. In this paper, the order of events is put back into the data model to provide topological depictions of the dynamics that are embodied in patients’ movement across a complex healthcare system. Result: The results show that directed graphs are well suited to the task of depicting the way that the diverse components of the system are functionally coupled—or remain disconnected—by patient journeys. Conclusion: By setting the resolution on the graph topology visualization, important characteristics can be highlighted, including highly prevalent repeating sequences of service events readily interpretable by clinical subject matter experts. Moreover, this methodology provides a first step in addressing the challenge of locating potential operational problems for patients with complex issues engaging with a complex healthcare service system.
Full article
(This article belongs to the Special Issue Feature Papers in Clinical Informatics Section)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Utilizing Generative Adversarial Networks for Acne Dataset Generation in Dermatology
by
Aravinthan Sankar, Kunal Chaturvedi, Al-Akhir Nayan, Mohammad Hesam Hesamian, Ali Braytee and Mukesh Prasad
BioMedInformatics 2024, 4(2), 1059-1070; https://doi.org/10.3390/biomedinformatics4020059 - 9 Apr 2024
Abstract
Background: In recent years, computer-aided diagnosis for skin conditions has made significant strides, primarily driven by artificial intelligence (AI) solutions. However, despite this progress, the efficiency of AI-enabled systems remains hindered by the scarcity of high-quality and large-scale datasets, primarily due to privacy
[...] Read more.
Background: In recent years, computer-aided diagnosis for skin conditions has made significant strides, primarily driven by artificial intelligence (AI) solutions. However, despite this progress, the efficiency of AI-enabled systems remains hindered by the scarcity of high-quality and large-scale datasets, primarily due to privacy concerns. Methods: This research circumvents privacy issues associated with real-world acne datasets by creating a synthetic dataset of human faces with varying acne severity levels (mild, moderate, and severe) using Generative Adversarial Networks (GANs). Further, three object detection models—YOLOv5, YOLOv8, and Detectron2—are used to evaluate the efficacy of the augmented dataset for detecting acne. Results: Integrating StyleGAN with these models, the results demonstrate the mean average precision (mAP) scores: YOLOv5: 73.5%, YOLOv8: 73.6%, and Detectron2: 37.7%. These scores surpass the mAP achieved without GANs. Conclusions: This study underscores the effectiveness of GANs in generating synthetic facial acne images and emphasizes the importance of utilizing GANs and convolutional neural network (CNN) models for accurate acne detection.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
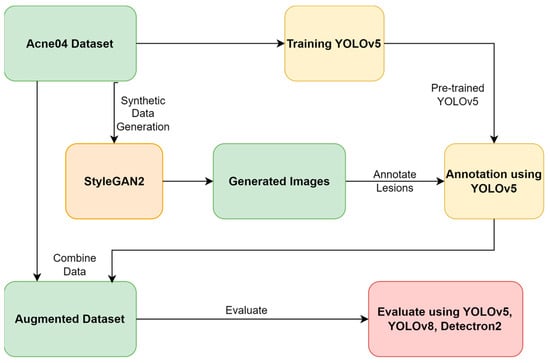
Figure 1
Open AccessArticle
A Comprehensive Analysis of Trapezius Muscle EMG Activity in Relation to Stress and Meditation
by
Mohammad Ahmed, Michael Grillo, Amirtaha Taebi, Mehmet Kaya and Peshala Thibbotuwawa Gamage
BioMedInformatics 2024, 4(2), 1047-1058; https://doi.org/10.3390/biomedinformatics4020058 - 9 Apr 2024
Abstract
Introduction: This study analyzes the efficacy of trapezius muscle electromyography (EMG) in discerning mental states, namely stress and meditation. Methods: Fifteen healthy participants were monitored to assess their physiological responses to mental stressors and meditation. Sensors were affixed to both the right and
[...] Read more.
Introduction: This study analyzes the efficacy of trapezius muscle electromyography (EMG) in discerning mental states, namely stress and meditation. Methods: Fifteen healthy participants were monitored to assess their physiological responses to mental stressors and meditation. Sensors were affixed to both the right and left trapezius muscles to capture EMG signals, while simultaneous electroencephalography (EEG) was conducted to validate cognitive states. Results: Our analysis of various EMG features, considering frequency ranges and sensor positioning, revealed significant changes in trapezius muscle activity during stress and meditation. Notably, low-frequency EMG features facilitated enhanced stress detection. For accurate stress identification, sensor configurations can be limited to the right trapezius muscle. Furthermore, the introduction of a novel method for determining asymmetry in EMG features suggests that applying sensors on bilateral trapezius muscles can improve the detection of mental states. Conclusion: This research presents a promising avenue for efficient cognitive state monitoring through compact and convenient sensing.
Full article
(This article belongs to the Special Issue Editor's Choices Series for Clinical Informatics Section)
►▼
Show Figures
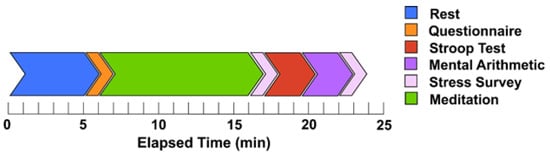
Figure 1
Open AccessArticle
Quantifying Inhaled Concentrations of Particulate Matter, Carbon Dioxide, Nitrogen Dioxide, and Nitric Oxide Using Observed Biometric Responses with Machine Learning
by
Shisir Ruwali, Shawhin Talebi, Ashen Fernando, Lakitha O. H. Wijeratne, John Waczak, Prabuddha M. H. Dewage, David J. Lary, John Sadler, Tatiana Lary, Matthew Lary and Adam Aker
BioMedInformatics 2024, 4(2), 1019-1046; https://doi.org/10.3390/biomedinformatics4020057 - 3 Apr 2024
Abstract
Introduction: Air pollution has numerous impacts on human health on a variety of time scales. Pollutants such as particulate matter—PM1 and PM2.5, carbon dioxide (CO2), nitrogen dioxide (NO2), and nitric oxide (NO) are exemplars of the
[...] Read more.
Introduction: Air pollution has numerous impacts on human health on a variety of time scales. Pollutants such as particulate matter—PM1 and PM2.5, carbon dioxide (CO2), nitrogen dioxide (NO2), and nitric oxide (NO) are exemplars of the wider human exposome. In this study, we adopted a unique approach by utilizing the responses of human autonomic systems to gauge the abundance of pollutants in inhaled air. Objective: To investigate how the human body autonomically responds to inhaled pollutants in microenvironments, including PM1, PM2.5, CO2, NO2, and NO, on small temporal and spatial scales by making use of biometric observations of the human autonomic response. To test the accuracy in predicting the concentrations of these pollutants using biological measurements of the participants. Methodology: Two experimental approaches having a similar methodology that employs a biometric suite to capture the physiological responses of cyclists were compared, and multiple sensors were used to measure the pollutants in the air surrounding them. Machine learning algorithms were used to estimate the levels of these pollutants and decipher the body’s automatic reactions to them. Results: We observed high precision in predicting PM1, PM2.5, and CO2 using a limited set of biometrics measured from the participants, as indicated with the coefficient of determination (R2) between the estimated and true values of these pollutants of 0.99, 0.96, and 0.98, respectively. Although the predictions for NO2 and NO were reliable at lower concentrations, which was observed qualitatively, the precision varied throughout the data range. Skin temperature, heart rate, and respiration rate were the common physiological responses that were the most influential in predicting the concentration of these pollutants. Conclusion: Biometric measurements can be used to estimate air quality components such as PM1, PM2.5, and CO2 with high degrees of accuracy and can also be used to decipher the effect of these pollutants on the human body using machine learning techniques. The results for NO2 and NO suggest a requirement to improve our models with more comprehensive data collection or advanced machine learning techniques to improve the results for these two pollutants.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
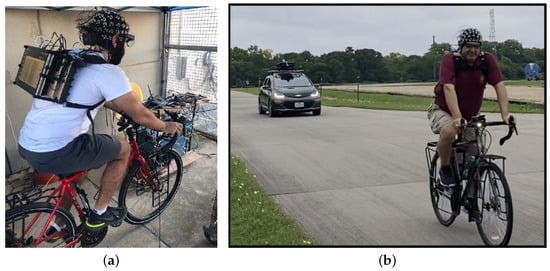

Figure 1
Open AccessArticle
Design and Modelling of an Induction Heating Coil to Investigate the Thermal Response of Magnetic Nanoparticles for Hyperthermia Applications
by
Philip Drake, Ali Algaddafi, Thomas Swift and Raed A. Abd-Alhameed
BioMedInformatics 2024, 4(2), 1006-1018; https://doi.org/10.3390/biomedinformatics4020056 - 2 Apr 2024
Abstract
Magnetic Field Hyperthermia is a technique where tumours are treated through an increase in local temperature upon exposure to alternating magnetic fields (AMFs) that are mediated by magnetic nano-particles (MNPs). In an AMF, these particles heat-up and kill the cells. The relationship between
[...] Read more.
Magnetic Field Hyperthermia is a technique where tumours are treated through an increase in local temperature upon exposure to alternating magnetic fields (AMFs) that are mediated by magnetic nano-particles (MNPs). In an AMF, these particles heat-up and kill the cells. The relationship between an AMF and the heating-rate is complex, leading to confusion when comparing data for different MNP and AMF conditions. This work allows for the thermal-response to be monitored at multiple AMF amplitudes while keeping other parameters constant. An induction-heating coil was designed based on a Zero-Voltage-Zero-Current (ZVZC) resonant circuit. The coil operates at 93 kHz with a variable DC drive-voltage (12–30 V). NEC4 software was used to model the magnetic field distribution, and MNPs were synthesised by the coprecipitation method. The magnetic field was found to be uniform at the centre of the coil and ranged from 1 kAm−1 to 12 kAm−1, depending on the DC drive-voltage. The MNPs were found to have a specific absorption rate (SAR) of 1.37 Wg−1[Fe] and 6.13 Wg−1[Fe] at 93 kHz and 2.1 kAm−1 and 12.6 kAm−1, respectively. The measured SAR value was found to be directly proportional to the product of the frequency and field-strength (SARα f Ho). This leads to the recommendation that, when comparing data from various groups, the SAR value should be normalized following this relationship and not using the more common relationship based on the square of the field intensity (SARα f Ho2).
Full article
(This article belongs to the Special Issue Feature Papers in Clinical Informatics Section)
►▼
Show Figures
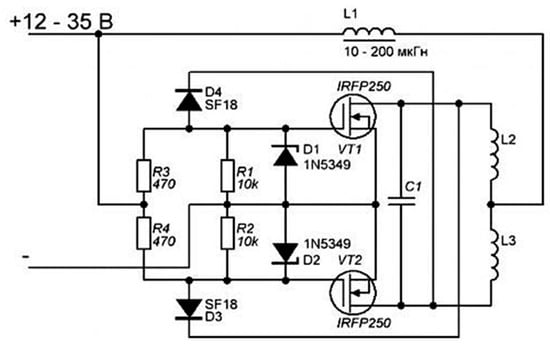
Figure 1
Open AccessArticle
Advancing Early Detection of Breast Cancer: A User-Friendly Convolutional Neural Network Automation System
by
Annie Dequit and Fatema Nafa
BioMedInformatics 2024, 4(2), 992-1005; https://doi.org/10.3390/biomedinformatics4020055 - 1 Apr 2024
Abstract
Background: Deep learning models have shown potential in improving cancer diagnosis and treatment. This study aimed to develop a convolutional neural network (CNN) model to predict Invasive Ductal Carcinoma (IDC), a common type of breast cancer. Additionally, a user-friendly interface was designed to
[...] Read more.
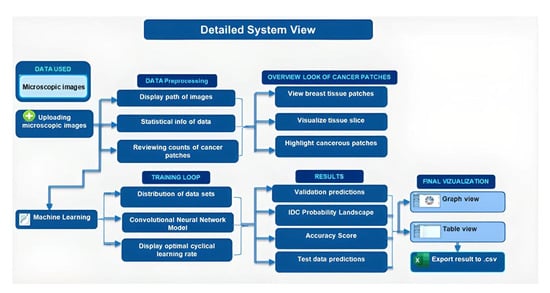
Background: Deep learning models have shown potential in improving cancer diagnosis and treatment. This study aimed to develop a convolutional neural network (CNN) model to predict Invasive Ductal Carcinoma (IDC), a common type of breast cancer. Additionally, a user-friendly interface was designed to facilitate the use of the model by healthcare professionals. Methods: The CNN model was trained and tested using a dataset of high-resolution microscopic images derived from 162 whole-mount slide images of breast cancer specimens. These images were meticulously scanned at 40× magnification using a state-of-the-art digital slide scanner to capture detailed information. Each image was then divided into 277,524 patches of 50 × 50 pixels, resulting in a diverse dataset containing 198,738 IDC-negative and 78,786 IDC-positive patches. Results: The model achieved an accuracy of 98.24% in distinguishing between benign and malignant cases, demonstrating its effectiveness in cancer detection. Conclusions: This study suggests that the developed CNN model has promising potential for clinical applications in breast cancer diagnosis and personalized treatment strategies. Our study further emphasizes the importance of accurate and reliable cancer detection methods for timely diagnosis and treatment. This study establishes a foundation for utilizing deep learning models in future cancer treatment research by demonstrating their effectiveness in analyzing large and complex datasets. This approach opens exciting avenues for further research and potentially improves our understanding of cancer and its treatment.
Full article
(This article belongs to the Special Issue Feature Papers in Clinical Informatics Section)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Advancing Early Leukemia Diagnostics: A Comprehensive Study Incorporating Image Processing and Transfer Learning
by
Rezaul Haque, Abdullah Al Sakib, Md Forhad Hossain, Fahadul Islam, Ferdaus Ibne Aziz, Md Redwan Ahmed, Somasundar Kannan, Ali Rohan and Md Junayed Hasan
BioMedInformatics 2024, 4(2), 966-991; https://doi.org/10.3390/biomedinformatics4020054 - 1 Apr 2024
Abstract
Disease recognition has been revolutionized by autonomous systems in the rapidly developing field of medical technology. A crucial aspect of diagnosis involves the visual assessment and enumeration of white blood cells in microscopic peripheral blood smears. This practice yields invaluable insights into a
[...] Read more.
Disease recognition has been revolutionized by autonomous systems in the rapidly developing field of medical technology. A crucial aspect of diagnosis involves the visual assessment and enumeration of white blood cells in microscopic peripheral blood smears. This practice yields invaluable insights into a patient’s health, enabling the identification of conditions of blood malignancies such as leukemia. Early identification of leukemia subtypes is paramount for tailoring appropriate therapeutic interventions and enhancing patient survival rates. However, traditional diagnostic techniques, which depend on visual assessment, are arbitrary, laborious, and prone to errors. The advent of ML technologies offers a promising avenue for more accurate and efficient leukemia classification. In this study, we introduced a novel approach to leukemia classification by integrating advanced image processing, diverse dataset utilization, and sophisticated feature extraction techniques, coupled with the development of TL models. Focused on improving accuracy of previous studies, our approach utilized Kaggle datasets for binary and multiclass classifications. Extensive image processing involved a novel LoGMH method, complemented by diverse augmentation techniques. Feature extraction employed DCNN, with subsequent utilization of extracted features to train various ML and TL models. Rigorous evaluation using traditional metrics revealed Inception-ResNet’s superior performance, surpassing other models with F1 scores of 96.07% and 95.89% for binary and multiclass classification, respectively. Our results notably surpass previous research, particularly in cases involving a higher number of classes. These findings promise to influence clinical decision support systems, guide future research, and potentially revolutionize cancer diagnostics beyond leukemia, impacting broader medical imaging and oncology domains.
Full article
(This article belongs to the Special Issue Advances in Quantitative Imaging Analysis: From Theory to Practice)
►▼
Show Figures
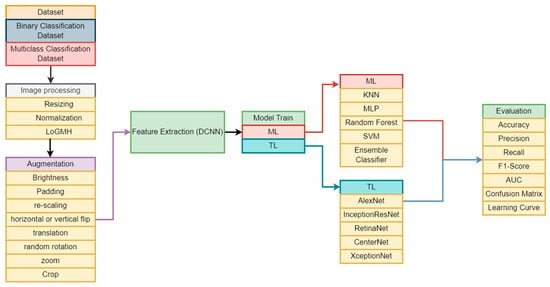
Figure 1
Open AccessArticle
A Methodological Approach to Extracting Patterns of Service Utilization from a Cross-Continuum High Dimensional Healthcare Dataset to Support Care Delivery Optimization for Patients with Complex Problems
by
Jonas Bambi, Yudi Santoso, Hanieh Sadri, Ken Moselle, Abraham Rudnick, Stan Robertson, Ernie Chang, Alex Kuo, Joseph Howie, Gracia Yunruo Dong, Kehinde Olobatuyi, Mahdi Hajiabadi and Ashlin Richardson
BioMedInformatics 2024, 4(2), 946-965; https://doi.org/10.3390/biomedinformatics4020053 - 1 Apr 2024
Cited by 1
Abstract
Background: Optimizing care for patients with complex problems entails the integration of clinically appropriate problem-specific clinical protocols, and the optimization of service-system-encompassing clinical pathways. However, alignment of service system operations with Clinical Practice Guidelines (CPGs) is far more challenging than the time-bounded alignment
[...] Read more.
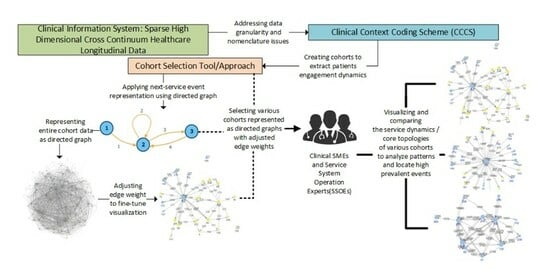
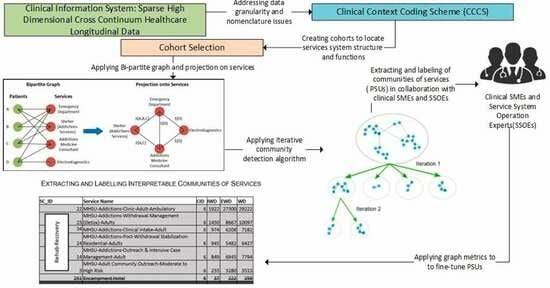
Background: Optimizing care for patients with complex problems entails the integration of clinically appropriate problem-specific clinical protocols, and the optimization of service-system-encompassing clinical pathways. However, alignment of service system operations with Clinical Practice Guidelines (CPGs) is far more challenging than the time-bounded alignment of procedures with protocols. This is due to the challenge of identifying longitudinal patterns of service utilization in the cross-continuum data to assess adherence to the CPGs. Method: This paper proposes a new methodology for identifying patients’ patterns of service utilization (PSUs) within sparse high-dimensional cross-continuum health datasets using graph community detection. Result: The result has shown that by using iterative graph community detections, and graph metrics combined with input from clinical and operational subject matter experts, it is possible to extract meaningful functionally integrated PSUs. Conclusions: This introduces the possibility of influencing the reorganization of some services to provide better care for patients with complex problems. Additionally, this introduces a novel analytical framework relying on patients’ service pathways as a foundation to generate the basic entities required to evaluate conformance of interventions to cohort-specific clinical practice guidelines, which will be further explored in our future research.
Full article
(This article belongs to the Special Issue Feature Papers in Clinical Informatics Section)
►▼
Show Figures
Graphical abstract
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
BioMedInformatics, Cancers, Current Oncology, Hemato, Hematology Reports
Myeloma and Leukemia-Challenges and Current Treatment Options
Topic Editors: Giovanni Martinelli, Claudio CerchioneDeadline: 20 June 2024
Topic in
Algorithms, BDCC, BioMedInformatics, Information, Mathematics
Machine Learning Empowered Drug Screen
Topic Editors: Teng Zhou, Jiaqi Wang, Youyi SongDeadline: 31 August 2024
Topic in
Biology, BioMedInformatics, Cancers, Genes, IJMS
The 22nd International Conference on Bioinformatics (InCoB 2023): Translational Bioinformatics Transforming Life
Topic Editors: Jyotsna Batra, Srilakshmi Srinivasan, Shoba Ranganathan, Asif M. Khan, Harpreet SinghDeadline: 15 November 2024
Topic in
Biology, Cancers, Molecules, IJMS, BioMedInformatics, Current Oncology
Learning Machines and Drug Discovery: A New Era in Cancer
Topic Editors: Satwinderjeet Kaur, Atiah H. AlmalkiDeadline: 20 December 2024

Conferences
Special Issues
Special Issue in
BioMedInformatics
Features of Bioinformatic Analyses for SARS-CoV-2 Infections and Vaccination
Guest Editor: Yasunari MatsuzakaDeadline: 30 June 2024
Special Issue in
BioMedInformatics
Advances in Quantitative Imaging Analysis: From Theory to Practice
Guest Editors: Federico Mastroleo, Angela Ammirabile, Giulia MarvasoDeadline: 30 November 2024
Special Issue in
BioMedInformatics
Advances in Structural Bioinformatics and Next-Generation Sequence Analysis for Drug Design
Guest Editor: Alexandre G. De BrevernDeadline: 31 December 2024






